SARS-CoV-2 and the Role of Orofecal Transmission: Evidence Brief
July 16, 2020
Jefferson T, Spencer EA, Brassey J, Heneghan C.
In: Analysis of the Transmission Dynamics of COVID-19: An Open Evidence Review.
Published Online July 17, 2020.
http://www.cebm.net/evidence-synthesis/transmission-dynamics-of-covid-19/
Evidence Brief pdf
Verdict
Various observational and mechanistic evidence presented throughout this evidence brief, support the hypothesis that SARS-CoV-2 can infect and be shed from the human gastrointestinal tract.
Policy
Policy should emphasise routine surveillance of food, wastewaters and effluent. The importance of strict personal hygiene measures, chlorine-based disinfection of surfaces in locations with presumed or known SARS CoV-2 activity should form part of public policy and education campaigns. Stool testing should be carried out in dischargees from the hospital or other holding facilities well before discharge date and discharge should be conditional either on cessation of fecal excretion or strict quarantine and personal hygiene measures in those still excreting viral particles by stool independently from respiratory excretion.
Further research
Each outbreak should be investigated and a report be made publicly available rapidly. Testing of stools should be carried out in all people involved in the outbreak.
As there is coherent evidence of ingestion, penetration of enterocytes and excretion of live SARs CoV-2 in possible analogy with SARs and MERS agents we believe this working hypothesis should be tested by conducting case-control studies during the investigation of outbreaks following a set protocol.
Cases would be cases of Covid-19 (with a subset by symptom presence and severity) either fecally excreting virions or not (cases and contacts) and controls would be healthy matches.
Exposure to potentially fecally contaminated materials and protective measures taken would be elicited at interview. To minimise the play of recall and ascertainment bias, interviewers should be blind to fecal excretion status and the interview should take place as soon as possible after the event.
Viability of fecal isolates and their possible pathogenicity should be tested in outbreaks, irrespective of the presence of symptoms or nasal swab positivity.
Background
Understanding how, when and in what types of settings SARS-CoV-2 spreads between people is critical to developing effective public health and infection prevention measures to break chains of transmission.1
Current evidence suggests SARS-CoV-2 is primarily transmitted via respiratory droplets and contact routes and can occur between pre-symptomatic or symptomatic infected individuals to others in close contact.
SARS-CoV-2 has been shown to contaminate and survive on certain surfaces, but currently, no reports have directly demonstrated fomite to human transmission. SARS-CoV-2 has also been detected in the feces of some patients which taken together with fomite transmission suggest the possibility that SARS-CoV-2 could transmit via the orofecal route.
Orofecal describes a route of transmission where the virus in fecal particles can pass from one person to the mouth of another. Main causes include lack of adequate sanitation and poor hygiene practices. Fecal contamination of food is another form of orofecal transmission.
Other single-stranded RNA viruses such as noroviruses are transmitted primarily through the orofecal route, either by consumption of contaminated food or water or from direct person to person to person spread.
Methods
We are undertaking an open evidence review investigating factors and circumstances that impact on the transmission of SARS-CoV-2, based on our published protocol. In brief, this review aims to identify and evaluate relevant articles (peer-reviewed or awaiting peer review) that examine the mode of viral transmission and ecological variables influencing the mode of transmission. Studies with modelling are only included if they report transmission outcome data, not predicted outcomes. We assess study quality based on five criteria and report important findings on an ongoing basis. When necessary we wrote to several authors of included studies for further details or clarification on the content of their articles.
Evidence update
In this first version we have summarized 36 studies examining the potential role of orofecal transmission of SARS-CoV-2 (see Table of Included Studies. COVID-19 transmission dynamics.), and included mechanistic and observational evidence from a further 22 studies (see References. Orofecal transmission potential of COVID-19). Overall the evidence is low to moderate quality (see Quality of Included Studies. COVID-19 transmission dynamics.). We provide a narrative summary of the evidence through a link at the end of each title in the left-hand column. The original paper is accessible through our summary.
SARS-COV-2 excretion and detection via the orofecal route
A review of 29 relevant studies (23 published and 6 preprints) reported that approximately 12% of patients with SARS-CoV-2 infection reported gastrointestinal symptoms, including diarrhoea, nausea, and vomiting. Eight included studies reported the detection of viral RNA of SARS-CoV-2 in stool and the pooled analysis reported that RNA shedding in stool was detected in up to 41% of COVID-19 patients [Parasa 2020].
In a group of 206 hospitalised patients with mild COVID-19, digestive symptoms were present in over half: 48 presented with a digestive symptom alone, 69 with both digestive and respiratory symptoms. [Han C 2020] Patients with digestive symptoms had a longer duration between symptom onset and viral clearance and were more likely to be fecal virus-positive (73% vs 14%, P=0.033) than those with respiratory symptoms. A further review of fourteen small studies that vary from 153 cases to a single case report reported that SARS-CoV-2 may be transmitted oro-fecally [Amirian 2020]
A second review of observational and mechanistic evidence provides further support that SARS-CoV-2 can infect and be shed from the human gastrointestinal tract. [Ding S 2020]
A review of 26 studies reported that there is a high rate of PCR positive persistence of SARS-CoV-2 in fecal samples of patients with COVID-19. [Gupta 2020] The review included 824 patients across the studies and 540 tested for faecal viral RNA: 291 (54%) had positive faecal RT-PCR tests. None of the studies was designed to detect live fecal virus except for the study by [Wang 2020].
Tian 2020 reviewed 15 included studies with data from 2,023 patients (mainly from China) and showed that gastrointestinal symptoms are common in COVID-19 patients and were observed with increased prevalence as the epidemic progressed in China.
Timing of fecal shedding
A retrospective study of 133 hospitalised COVID-19 patients identified 22 whose sputum or fecal samples tested positive, after their pharyngeal swabs became negative. [Chen 2020] This study assessed the results of real-time quantitative fluorescence polymerase chain reaction (RT-qPCR) for SARS–CoV2 RNA of sputum and fecal samples from a group of hospitalised COVID-19 patients, after conversion of their pharyngeal samples from positive to negative.
A convenience sample was retrospectively identified, of patients admitted to Beijing Ditan Hospital, Capital Medical University, with a diagnosis of COVID-19 and paired RT-qPCR testing of pharyngeal swabs with either sputum or feces samples. Among 133 patients admitted with COVID-19 from 20 January to 27 February 2020, 22 patients (4 of whom were children) with an initial or follow-up positive sputum or fecal samples paired with a follow-up negative pharyngeal sample were identified. From these 22 patients, 545 samples were available, including 209 pharyngeal swabs, 262 sputum samples, and 74 feces samples. Chen 2020 does not report on the viability of the viral fecal samples.
Cheung 2020 reported in a cohort of 59 patients in Hong Kong with COVID-19 that fecal discharge continues long after respiratory shedding of COVID-19 has ceased. Gupta 2020 reported that the duration for faecal shedding of viral RNA after clearance of respiratory samples ranged from 1 to 33 days and in one patient up to 47 days from symptom onset.
In the review by Tian 2020 fecal PCR testing was shown to be as accurate as respiratory specimen PCR detection, and fecal excretion persisted after sputum excretion in 23% of patients for 1 to 11 days. A systematic review of four case series including 36 children with mild COVID-19 found that fecal shedding occurs on average nine days longer than by the respiratory route. Three times as many children had SARS-CoV-2 shedding in stools after 14 days of symptoms onset compared to respiratory samples, RR= 3.2 (95%CI 1.2 to 8.9). Because of the delayed fecal shedding. [Santos 2020] The review authors were unsure whether the fecally shed viruses were viable and hence infectious.
A short review by Donà 2020 including six studies of children reports the orofecal route is an alternative route of transmission, regardless of presenting COVID-19 symptomatology. Citing evidence from the SARS-CoV-1 outbreak that fecal excretion could be ongoing even after 30 days from symptom onset, the review authors recommend that exclusion of SARS-CoV-2 infection by single time point nasopharyngeal swabs should not be used in children.
Live SARS-COV-2 excretion by the fecal route
Many studies have identified SARS-CoV-2 RNA in fecal samples. Most studies have been unable to identify live viruses within fecal samples. Three case studies (Wang W 2020, Xiao 2020b and Yong Z 2020) report isolating the virus (in 2 patients, 1 patient and 1 patient respectively).
Wang 2020 detected SARS-CoV-2 in the feces of 44 of 153 (29%) specimens collected from 205 patients with COVID-19 from three hospitals in the Hubei and Shandong provinces and Beijing. RNA was extracted by RT-PCR targeting the open reading frame 1ab gene of SARS-CoV-2. A cycle threshold value less than 40 was interpreted as positive for SARS-CoV-2 RNA. Four positive fecal specimens with high copy numbers were then cultured, and electron microscopy performed to detect live virus. Live SARS-CoV-2 virus was observed in the stool sample of two patients who did not have diarrhea. The Parasa 2020 review interpreted this as evidence that the gastrointestinal tract supports the growth of SARS-CoV-2 to an extent similar to previous SARS infections, and it might explain some of the rapid spread of disease.
Xiao 2020b Identified by Susan Amirian undertook a case-series of 28 hospitalised patients with severe COVID-19 for whom faeces samples were available.
Among the specimens collected 12 were positive for viral RNA at least one-time point. SARS-CoV-2 virus was successfully isolated from two of the viral RNA–positive patients. Viral particles that were visible were spherical and had distinct surface spike protein projections, consistent with a previously published SARS-CoV2 image. Four serial feces samples from a seriously ill 78-year old patient with COVID-19 all tested positive for viral RNA; the patient subsequently died. Viral antigen was also detected in gastrointestinal epithelial cells of a biopsy sample, from this 78-year old COVID-19 patient.
Yong Z 2020 isolated live virus from the stools of one severe pneumonia case, pointing to a possible orofecal spread. There were unclear and sparse clinical details in the study report.
A preprint identified by Susan Amirian on the patient-derived mutations impact pathogenicity of SARS-CoV-2 seems to indicate that 3/3 stools contained live virus (see pdf p 24)2 and is capable of replicating in stool samples.
Tian 2020 review (mainly based on Chinese data) reported that angiotensin-converting enzyme 2 (ACE2) and virus nucleocapsid protein were detected in gastrointestinal epithelial cells, and infectious virus particles were isolated from faeces. A case series of 28 hospitalised patients for whom feces samples were collected reported that SARS-CoV-2 virus was successfully isolated from two of the viral RNA–positive patients. Viral particles that were visible were spherical and had distinct surface spike protein projections, consistent with a previously published SARS-CoV-2 image. [Xiao F 2020 b] Four serial feces samples from a seriously ill 78-year old patient with COVID-19 all tested positive for viral RNA; the patient subsequently died. Viral antigen was also detected in gastrointestinal epithelial cells of a biopsy sample, from this 78-year old COVID-19 patient.
Zang R 2020 undertook a laboratory study that reported that human enterocytes express high ACE2 receptor levels, which could support infection with SARS-CoV-2. However, they reported the virus is rapidly inactivated in the GI tract. This poses the question of how enveloped coronaviruses could survive the low pH of the stomach and the effect of bile salts in the small bowel.
An assessment of viral RNA in feces from 71 hospitalized patients with SARS-CoV-2 reported that viral RNA and viral nucleocapsid protein in gastrointestinal tissues was extracted from one patient. [Xiao F 2020] From the 1st to 14th February 2020, clinical specimens, including serum, nasopharyngeal, and oropharyngeal swabs; urine; stool; and tissues were obtained from 73 hospitalized patients infected with SARS-CoV-2 were obtained and tested for SARS-CoV-2 RNA. Immunofluorescent data showed that ACE2 protein (a cell receptor for SARS-CoV-2) is expressed in the glandular cells of gastric, duodenal, and rectal epithelia, supporting the entry into the host cells. SARS-CoV-2 might therefore both infect the gastrointestinal system and transmit via the orofecal route, and detection of viral nucleocapsid protein in rectal epithelial cells suggests that some infectious viral particles can survive the adverse gastrointestinal environment.
Ding S 2020 review cited evidence that SARS-CoV-2 can survive the adverse conditions in the gastrointestinal system. Both the S glycoproteins and the viral envelope must resist degradation to infect enterocytes. Heavy glycosylation of the large spike S protein has been shown to lead to resistance to the proteases, the low pH and bile salts found in the gastrointestinal system. Some gastric processes may actually facilitate viral entry into the enterocytes: in bovine coronavirus, one specific site on the S glycoprotein has to be cleaved by an intracellular protease or trypsin to activate viral infectivity and cell fusion.3
Cruise Ships
Evidence of fomite transmission of COVID-19 first emerged due to quarantine on the Diamond Princess Cruise Ship. The infection rate onboard was high and clinical harm was considerable onboard the ship. [Yamahata Y] Of those tested, 696 (19%) tested positive for COVID-19, of which 410 (59%) were asymptomatic. The structure of the cruise ship made it difficult to carry out the medical services required for an outbreak of an emerging infectious disease. The authors noted that one of the elevator halls may have been a transmission point because it was impossible for infected and non-infected people to use the elevator separately. Since all passengers were isolated in each cabin, supplies had to be delivered daily to each room. The disposal of the sewage was also a challenge and eventually, a service tender came to extract sewage from the ship. Fifteen of the 20 confirmed cases occurred in crew members among the food service workers, and sixteen cases occurred among those with cabins on deck 3, the deck on which the food service workers lived. [Kakimoto 2020]
Choir Practice
The high SARS-CoV-2 attack rate following exposure at a choir practice in Skagit County, Washington suggested that transmission of COVID-19 was facilitated by close proximity and physical contact and may have been augmented by the act of singing. [Hamner L 2020]. The 2.5-hour singing practice, however, also provided opportunities for orofecal transmission, as members shared snacks at the end of the practice.
Acute Healthcare Settings
An investigation of SARS-CoV-2 on surfaces and air contamination in an acute healthcare setting in London found that many hospital surfaces and air samples contained viral RNA. [Zhou J] Viral RNA was detected on 114/218 (52%) of surfaces and 14/31 (30%) of air samples. The proportion of surface samples contaminated with viral RNA varied by item sampled and by clinical area. And although viral RNA was more likely to be found in areas immediately occupied by COVID-19 patients than in other areas, 45% of the samples were from areas not immediately occupied by a COVID-19 patient. Surface contamination was detected on a range of items: computer keyboards, chairs, and alcohol dispensers had the highest proportion of samples. Viable virus, however, was not cultured from any of the air or surface samples.
An analysis of 626 surface swabs within the Zhongnan Medical Center in Wuhan between 7th February and 27th February 2020 found 14% of the 626 surface samples were positive for viral RNA. One third (32%) of the contaminated zones were found in the intensive care unit, 28% were in the obstetric ward specialized for pregnant women with COVID-19 and 20% were in the ward for COVID-19 patients. The most contaminated surfaces were self-service printers (20%), desktop/keyboard (17%), and door handles (16%). Hand sanitizer dispensers (20%) and gloves (15%) were the most contaminated protective equipment.4
From January 24th to February 4th, three infected patients in airborne infection isolation rooms with anterooms and bathrooms had surface environmental samples taken at 26 sites. In two symptomatic patients rooms, after routine cleaning all samples were negative. In the third patient’s room, samples were collected before routine cleaning and were found to be positive for 13 out of 15 (87%) sites (including air outlet fans) and 3 of 5 toilet sites (toilet bowl, sink, and door handle) positive. Patient C had upper respiratory tract involvement and two positive stool samples for SARS-CoV-2 on RT-PCR despite not having diarrhoea. 5
A study in February 2020 the infectious diseases hospital Nanjing, China randomly sampled the 3-bed isolation rooms of the COVID-19 designated infectious diseases hospital, Nanjing, China. [Ding Z 2020] Environmental sampling was also carried out in four isolation rooms, a nursing station, a corridor, an air-conditioning system and other spaces in the airborne infectious-disease zone on the fifth floor of the hospital. Sampling procedures are described accurately and the air sampler twice needed to be quarantined despite wearing full PPE. Airflow was also assessed between 4th and 5th floors in the building using a smoke tracer.
Of 107 surface samples (37 from toilets, 34 from other surfaces in isolation rooms and 36 from other surfaces outside isolation rooms). Four samples were positive (2 two ward door door-handles, one bathroom toilet toilet-seat cover and one bathroom door door-handle). Three were weakly positive from a bathroom toilet seat, one bathroom washbasin tap lever and one bathroom ceiling exhaust louvre. One of the 46 corridor air samples was weakly positive.
The paper has a very clear colour-coded figure showing the timeline of onset and hospitalisation dates and the sampling dates of the events in the isolation rooms containing beds 2 and 3, beds 16-18, beds 31 and 32, and bed 55, and their 10 patients. The sampling dates on which positive samples were detected are also shown by a red tick. In each room, a patient and his/her bed are shown in the same colour.
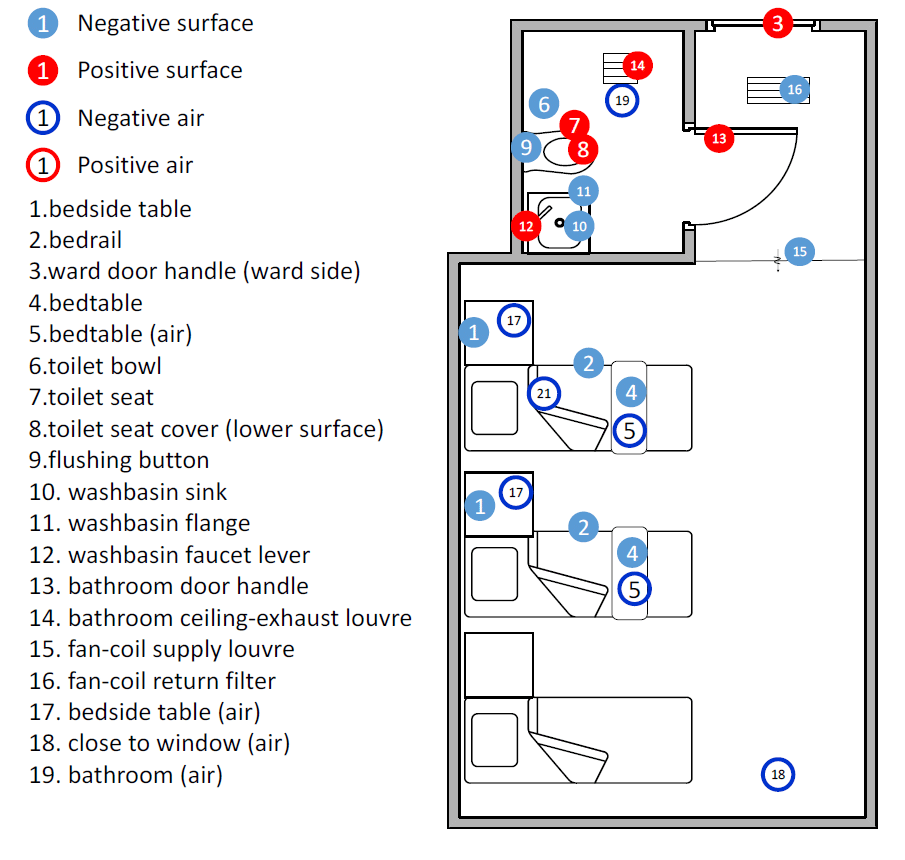
The figure shows a summary of the location of the samples taken in the isolation rooms. Reproduced from Ding Z, Qian H, Xu B, Huang Y, Miao T, Yen H-L, et al. Toilets dominate environmental detection of SARS-CoV-2 virus in a hospital. medRxiv 2020.04.03.20052175 2020
To determine the distribution of SAR-S-CoV-2 in hospital wards in Wuhan, China, Guo ZD6 tested air and surface samples. The virus was widely found on floors, computer mice, trash cans, and sickbed handrails. The nucleic acid test used did not indicate the amount of viable virus. The virus was also widely distributed in the air and patient masks also contained infected exhaled droplets and oral secretions. The authors recommended adequately disinfecting such masks before discarding them.
A study in the First Affiliated Hospital of Zhejiang University, China found that strict disinfection and hand hygiene could decrease the hospital-associated COVID-19 infection risk of the staff in isolation wards. The study monitored the presence of SARS-Cov-2 among hospital environment surfaces, sewage, and personal protective equipment (PPE) of staff in isolation wards in the First Affiliated Hospital of Zhejiang University, China. During the study period, 33 patients were hospitalized in isolation wards, and no SARS-Cov-2 RNA was detected among the 36 objects surface samples and nine staff PPE samples in isolation wards. 7
Surfaces of objects were wiped with 1000 mg/L chlorine-containing disinfectant every 4 hours in the isolation ICU ward and every 8 hours in general isolation wards. Preprocessing disinfection equipment was added before sewage drainage from the isolation wards into the final sewage disinfection pool. The sewage samples from the inlet pool were found to be positive, and from the outlet of the preprocessing pool weakly positive, but the sewage sample from the last disinfection pool was found to be negative. No viable virus was detected by culture.
Sewage
Concentration and detection of SARS coronavirus was found in sewage concentrates in China in two hospitals receiving SARS patients prior to disinfection, and occasionally after disinfection. There was no live SARS-CoV detected in the sewage in these assays. In this study, the authors found that the virus can survive for 14 days in sewage at 4°C, 2 days at 20°C, and its RNA could be detected for eight days although the virus had been inactivated. [Wang XV 2020]
Sentinel surveillance of SARS-CoV-2 in wastewater has been shown to anticipate the occurrence of COVID-19 cases. SARS-CoV-2 was detected in sewage 41 days before the declaration of the first COVID-19 case in Spain and in frozen samples dating back to 12 March 2019. If this is confirmed, the results suggest SARS CoV-2 has been around longer than first thought. [Chavarria-Miró G 2020]
Even at low COVID-19 prevalence, sewage surveillance could be a sensitive tool to monitor the viral circulation. SARS-CoV-2 RNA detection occurred in volumes of 250 mL of wastewaters collected in both areas at high (Milan) and low (Rome) epidemic circulation, according to clinical data. Six out of 12 samples were positive. [La Rosa G 2020] One of the positive results was obtained in a Milan wastewater sample collected three days after the first notified Italian case of autochthonous SARS-CoV-2 at Codogno. A second positive sample was taken on the 28th of February in Milan when COVID-19 cases were only 29.
Environmental screening may, therefore, be a very sensitive tool to gauge viral presence before clinical symptoms become apparent. SARS-CoV-2 was detected in the sewage of five sites a week after the first COVID-19 case in the Netherlands. The identification of viral antigens in sewage samples occurred when the observed COVID-19 prevalence was around or even below 1.0 case per 100,000 people. Stronger signals were observed when the prevalence was 3.5 cases per 100,000 people or more. [Medema G 2020] The authors did point out that none of the isolates are likely to be viable and the testing method needs refining.
A systematic review of what is known of the presence and survival of coronaviridae in various water settings included twelve articles. [La Rosa 2020] The authors remarked on the poor evidence base for enveloped viruses and that the methods of concentration and collection for these viridae may not be appropriate. Coroviridae have been isolated in different types of liquids from waste to surface water but in general, they appear to be unstable. Chlorination and higher temperatures lead to their inactivation. SARS-CoV was detected in wastewater, domestic sewage, and tap water for 2 days at 20°C and up to 14 days at 4°C. Chlorine was considered far more effective against CoV than other microorganisms because of its lytic action on the envelope. None of the samples for SARS-CoV 1 were viable but the presence of coronaviridae was widespread.
MERS-CoV and SARs-CoV-1
MERS-CoV has been shown to infect human primary intestinal epithelial cells, small intestine explants and intestinal organoids.8 MERS-CoV has been detected in 42% of milk samples collected from lactating camels where it can survive for a prolonged period. And in the largest hospital-associated MERS outbreak in the Republic of Korea in 2015 (n=186) a history of direct contact accounts for only one in ten cases.9 A study of human primary intestinal epithelial cells and small intestine explants of MERS-CoV patterns identified the viral replication intermediates in stool specimen. MERS-CoV was found to be resistant to fed-state gastrointestinal fluids but less tolerant to the high acidic fasted-state gastric fluid.
In 1977 the prolonged excretion of coronaviruses was first observed in feces.10 In the SARS-CoV-1 outbreak in 2002-03, a significant portion of patients had enteric involvement. For example, in the Toronto outbreak in 2003, 6% of 144 patients had diarrhoea on presentation.11 Also among 138 patients with SARS in Hong Kong, 20% presented with watery diarrhoea and 38% had symptoms of diarrhoea during the illness. Intestinal biopsy specimens showed the presence of active viral replication, and SARS-CoV RNA was detected in the stool of some patients for more than ten weeks after symptom onset.12 A retrospective study on specimens from 154 patients in Hong Kong with laboratory-confirmed SARS found the viral load to be the highest in stool specimens.13 Up to 70% of 75 patients in a community outbreak in Hong Kong developed watery diarrhea.14 This outbreak was linked to a faulty sewage system in the Amoy Gardens apartment complex, further suggesting orofecal transmission might be a route for transmission.15
The human gastrointestinal tract might be a primary infection site for SARS-CoV. Ding et al used a monoclonal antibody specific for the SARS‐CoV nucleoprotein, and probes for the RNA polymerase gene fragment in four patients who died from SARS‐CoV-1.16 Virus was detected in the stomach, small intestine, distal convoluted renal tubule, sweat gland, parathyroid, pituitary, pancreas, adrenal, liver and cerebrum. The authors discussed that viruses in contaminated food and water may enter the human body through epithelial cells covering the surface of the gastrointestinal tract, although there was no direct evidence to show that food‐borne transmission had occurred.
A study from the sewage of two hospitals receiving SARS patients in Beijing found no infectious SARS-CoV contamination in any of the samples collected, but did detect the nucleic acid in the sewage from the two hospitals before disinfection – providing further evidence that SARS-CoV-1 can be excreted by feces into the sewage system. 17
Orofecal transmission
Orofecal transmission occurs with a number of viruses. Enteroviruses, for example, are positive-sense single-stranded RNA viruses and transmit through the intestine. They affect millions worldwide each year and can be found in respiratory secretions and the stool of infected persons. Over 90% of infected individuals with enteroviruses have no symptoms or have non-specific symptoms. Norovirus is largely spread by the orofecal route through person-to-person contact, contaminated food or water, or from the aerosolized vomit of an infected person. It often occurs among those living in close quarters and leads to outbreaks on cruise ships and healthcare facilities that often requires quarantining of the facilities. Transmission of norovirus, therefore, shares some similarities with SARs-CoV-2 including superspreading events.
Mounting evidence suggests that superspreading events have driven many of the local epidemics. Subsequent to identifying the index case in a French holiday chalet, SARS-CoV-2 was detected in 11 additional people [Danis K 2020]; a single case travelling on a bus and attending a mass gathering appeared to be responsible for infecting 25 individuals [Chen Y 2020]; and the transmission of SARS-CoV-2 in South Korea was exacerbated by super spreading events in confined settings, including a hospital, a church and a gym. [Shim E 2020]
Cluster sizes of more than 100 cases have been identified for hospitals, elderly care, worker dormitories, food processing plants, prisons, schools, shopping, religious events and ships.18 The vast majority of these were indoor settings.
Some evidence suggests that close contact does not increase the risk of transmission. A convenience sample of 533 close contacts (404 actively monitored) of nine early travel-related cases in the United States identified two additional cases; both secondary cases were in spouses of travel-associated case-patients. [Burke 2020] No transmission was found among the 389 non-household contacts who completed active monitoring. For community contacts many reported having face-to-face contact (27/35; 77%) with the travel-associated case or spending time within 6 feet (34/38; 90%), and nearly all (43/45; 96%) could remember being in the same room as the travel-associated case-patient. Fewer (8/28; 29%) reported being within six feet of the patient while the patient was coughing. No community contacts were subsequently diagnosed with COVID-19.
While most human Coronaviruses are considered not to transmit fecally this is not the case in animals. Feline coronavirus, for instance, is typically shed in feces of healthy cats and transmitted by the orofecal route to other cats.19 Pigs are also infected by the transmissible gastroenteritis coronavirus via the fecal-oral route.20 Bat Coronavirus infects the gastrointestinal and respiratory tracts of bats seemingly without causing disease.21 Transmission following exposure to camel feces has also been considered to be biologically plausible, although no evidence indicates whether this is possible. 22
There is, however, evidence that SARS-CoV-2 can survive adverse conditions in the gastrointestinal system. It has been identified in endoscopic specimens of the esophagus, stomach, duodenum, and rectum of COVID-19 patients; substantial amounts of SARS-CoV-2 RNA have been consistently detected in stool specimens. [DIng S 2020] Various observational and mechanistic evidence presented throughout this evidence brief supports the hypothesis that SARS-CoV-2 can infect and be shed from the human gastrointestinal tract.
Acknowledgment
Drs Susan Amirian, Siyuan Ding and Sravanthi Parasato provided additional information for this brief.
Disclaimer: The article has not been peer-reviewed. The views expressed in this commentary represent the views of the authors and not necessarily those of the host institution, the NHS, the NIHR, or the Department of Health and Social Care. The views are not a substitute for professional medical advice. It will be regularly updated see the evidence explorer at https://www.cebm.net/evidence-synthesis/transmission-dynamics-of-covid-19/ for regular updates to the evidence summaries and briefs.
Authors:
Tom Jefferson, epidemiologist. Disclosure statement is here
Elizabeth Spencer is Epidemiology and Evidence Synthesis Researcher at the Centre for Evidence-Based Medicine.
Jon Brassey is the Director of Trip Database Ltd, Lead for Knowledge Mobilisation at Public Health Wales (NHS) and an Associate Editor at the BMJ Evidence-Based Medicine.
Carl Heneghan is Professor of Evidence-Based Medicine, Director of the Centre for Evidence-Based Medicine and Director of Studies for the Evidence-Based Health Care Programme. (Full bio and disclosure statement here)
This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License.
References
References. Orofecal transmission potential of COVID-19
- Transmission of SARS-CoV-2: implications for infection prevention precautions https://www.who.int/news-room/commentaries/detail/transmission-of-sars-cov-2-implications-for-infection-prevention-precautions
- Patient-derived mutations impact pathogenicity of SARS-CoV-2 Hangping Yao et al .https://www.medrxiv.org/content/10.1101/2020.04.14.20060160v2.full.pdf
- Holmes K.V. Enteric infections with coronaviruses and toroviruses. Novartis Found Symp. 2001;238:258–269. discussion 269–275.
- Ye G, Lin H, Chen S, et al. Environmental contamination of SARS-CoV-2 in healthcare
- Ong SWX, Tan YK, Chia PY, et al. Air, Surface Environmental, and Personal Protective Equipment Contamination by Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) From a Symptomatic Patient. JAMA. 2020;323(16):1610–1612. doi:10.1001/jama.2020.3227
- Guo ZD, Wang ZY, Zhang SF, et al. Aerosol and Surface Distribution of Severe Acute
- Wang J, Feng H, Zhang S, et al. SARS-CoV-2 RNA detection of hospital isolation wards
- Zhou J et al. Human intestinal tract serves as an alternative infection route for Middle East respiratory syndrome coronavirus. Sci Adv. 2017;3:eaao4966. https://advances.sciencemag.org/content/3/11/eaao4966
- Lee SS, Wong NS. Probable transmission chains of Middle East respiratory syndrome coronavirus and the multiple generations of secondary infection in South Korea. Int J Infect Dis. 2015 Sep; 38():65-7.
- Further studies on human enteric coronaviruses. Caul EO, Egglestone SI.
- Booth CM et al. Clinical features and short-term outcomes of 144 patients with SARS in the greater Toronto area. JAMA. 2003;289:2801–2809.
- Leung WK, To K-F, Chan PKS, Chan HLY, Wu AKL, Lee N, Yuen KY, Sung JJY. Enteric involvement of severe acute respiratory syndrome-associated coronavirus infection. Gastroenterology 125, 1011–1017 (2003).
- Hung IFN et al. Viral loads in clinical specimens and SARS manifestations. Emerg. Infect. Dis. 10, 1550–1557 (2004).
- Peiris JSM et al. Clinical progression and viral load in a community outbreak of coronavirus-associated SARS pneumonia: a prospective study. Lancet. 2003;361:1767–1772.
- WHO environmental health team reports on Amoy gardens. Available at: http://www.info.gov.hk/gia/general/200305/16/0516114.htm.
- Ding Y, He L, Zhang Q, et al. Organ distribution of severe acute respiratory syndrome (SARS) associated coronavirus (SARS-CoV) in SARS patients: implications for pathogenesis and virus transmission pathways. J Pathol. 2004;203(2):622-630. doi:10.1002/path.1560
- Wang XW, Li JS, Guo TK, et al. Concentration and detection of SARS coronavirus in sewage from Xiao Tang Shan Hospital and the 309th Hospital [published correction appears in J Virol Methods. 2005 Dec;130(1-2):210]. J Virol Methods. 2005;128(1-2):156-161. doi:10.1016/j.jviromet.2005.03.022
- Analysis of SARS-CoV-2 transmission clusters and superspreading events Gwen Knight, Quentin Leclerc and Adam Kucharski, on behalf of CMMID working group Report for SPI-M, 3rd June 2020. https://assets.publishing.service.gov.uk/government/uploads/system/uploads/attachment_data/file/897562/S0473_SPI-M_Superspreading_and_clusters.pdf
- Hartmann, Katrin (2005). “Feline infectious peritonitis”. Veterinary Clinics of North America: Small Animal Practice. 35 (1): 39–79. doi:10.1016/j.cvsm.2004.10.011
- Immune evasion of porcine enteric coronaviruses and viral modulation of antiviral innate signaling.
- Fan Y, Zhao K, Shi ZL, Zhou P. Bat Coronaviruses in China. Viruses. 2019;11(3):210. Published 2019 Mar 2. doi:10.3390/v11030210
- Killerby ME, Biggs HM, Midgley CM, Gerber SI, Watson JT. Middle East Respiratory Syndrome Coronavirus Transmission. Emerg Infect Dis. 2020;26(2):191-198. https://dx.doi.org/10.3201/eid2602.190697

